Exploring the Structural Biology of Cancer
12/13
Cells in our bodies have evolved to work together, constantly communicating with each other, sharing resources, and growing and dividing only when and where needed. Normal cells that become transformed into cancer cells go rogue and grow without this cooperation with other cells in the body. "Cancer" encompasses many different diseases, but cancer cells share similar characteristics, including ignoring normal controls on growth, evading the body's built-in defenses, tuning metabolism and their surrounding environment for faster growth, and in the most severe cases, invading other parts of the body. Cells typically gain these characteristics through changes in their genomes.
Structural biologists are studying all aspects of this process, including the mechanisms that underpin damage to and repair of the genome, the way that these genetic changes lead to cancer, and ways for us to use this knowledge to discover and develop new treatments to fight cancer.
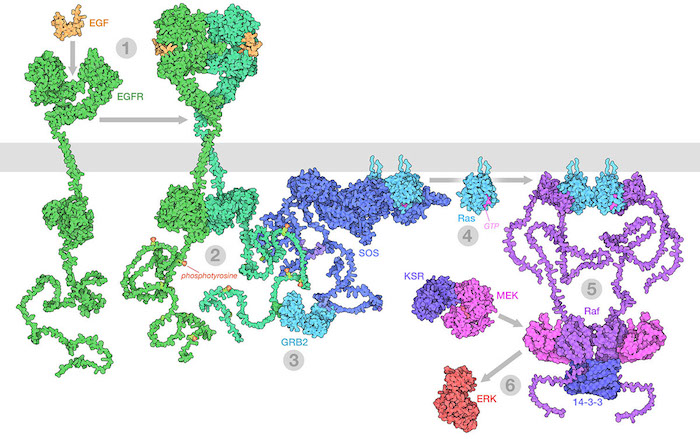 The Raf/MEK/ERK (MAPK) pathway is one of the ways that growth signals are disseminated inside the cell. 1) Epidermal growth factor (EGF) binds to its receptor (EGFR), causing it to dimerize. 2) The catalytic domains of EGFR phosphorylate tyrosines on the long unstructured EGFR tails. 3) GRB2 recognizes the phosphorylated tyrosines and recruits SOS. 4) SOS replaces GDP with GTP in Ras proteins. 5) The activated Ras forms nanoclusters that activate Raf, with the help of 14-3-3 protein. 6) Raf phosphorylates MEK, which is normally held in an inactive complex with KSR. Active MEK then phosphorylates ERK, a kinase that will activate many processes throughout the cell. Illustration created from decades of structural studies in PDB IDS 1egf, 1nql, 1m17, 2jwa, 3njp, 2gs6, 1gri, 1xd2, 3ksy, 5p21, 6xi7, 6q0j, 2y4i, 1pme, and unstructured chains from AlphaFold2.
The Raf/MEK/ERK (MAPK) pathway is one of the ways that growth signals are disseminated inside the cell. 1) Epidermal growth factor (EGF) binds to its receptor (EGFR), causing it to dimerize. 2) The catalytic domains of EGFR phosphorylate tyrosines on the long unstructured EGFR tails. 3) GRB2 recognizes the phosphorylated tyrosines and recruits SOS. 4) SOS replaces GDP with GTP in Ras proteins. 5) The activated Ras forms nanoclusters that activate Raf, with the help of 14-3-3 protein. 6) Raf phosphorylates MEK, which is normally held in an inactive complex with KSR. Active MEK then phosphorylates ERK, a kinase that will activate many processes throughout the cell. Illustration created from decades of structural studies in PDB IDS 1egf, 1nql, 1m17, 2jwa, 3njp, 2gs6, 1gri, 1xd2, 3ksy, 5p21, 6xi7, 6q0j, 2y4i, 1pme, and unstructured chains from AlphaFold2.Past news and events have been reported at the RCSB PDB website and past Newsletters.



